The diverse biological roles of the above examples highlight the broad importance of proteins that escort, facilitate or accelerate structural changes in non-coding RNA. Eukaryotic proteins with ATP-independent RNA chaperone properties include RNA recognition motif (RRM) proteins such as hnRNP A1 ( 27), viral proteins such as the well-studied retroviral nucleocapsid protein (NCp7) ( 28, 29), and La and Ro proteins ( 30, 31). H-NS and StpA, which interact with the bacterial nucleoid, also possess RNA chaperone activity ( 17). In bacteria, this type of passive RNA chaperone includes cold shock proteins (CSPs) ( 14), the Sm family protein Hfq ( 21), the FinO/ProQ family of RNA binding proteins ( 22), and ribosomal proteins S1 ( 23– 25) and S12 ( 26). This chapter, however, will focus on RNA binding proteins that “passively” remodel RNA structures without hydrolyzing ATP. Bacteria typically encode a handful (0–12) of DEAD-box proteins that mainly act in ribosome biogenesis and RNA turnover ( 12, 20). RNA helices can be actively unwound by DEAD-box proteins, which couple unfolding of the RNA structure to ATP hydrolysis ( 18, 19). RNA chaperones act by transiently binding and releasing RNA substrates, disrupting the RNA secondary and tertiary structure (unwinding and unfolding) or accelerating base pairing with a second RNA strand (annealing) ( 16, 17). coli ( 15), suggesting that such proteins broadly mitigate the effects of RNA misfolding. Moreover, over-expression of cold shock proteins and other RNA chaperones buffered deleterious mutations in E. 
For example, the up-regulation of cold shock domain RNA binding proteins during low temperature growth destabilizes RNA structures that would otherwise impair transcription elongation and translation initiation at low temperatures ( 14). These housekeeping and regulatory functions of RNA chaperones are particularly important at cold temperatures that hyperstabilize RNA structures. RNA chaperones also facilitate conformational rearrangements during ribosome biogenesis ( 12) and eukaryotic pre-mRNA splicing ( 13). These regulatory RNAs are chaperoned by diverse families of RNA binding proteins, and the loss of RNA chaperone proteins can lead to impaired growth, reduced tolerance to stress, and reduced virulence ( 6– 11). This is called its native state, and is the ground state of the system.Non-coding RNA sequences fold into useful structures that regulate gene expression as ribozymes, metabolite binding sensors, or antisense RNAs ( 1– 5). 
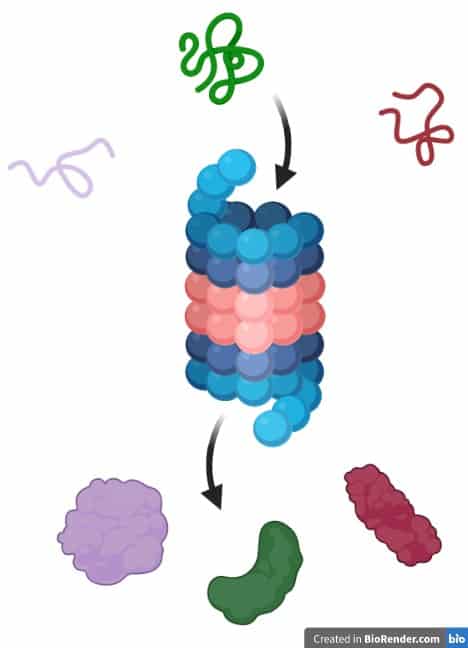
At the bottom of the funnel, the free energy is at a minimum and there is only one conformational state available to the protein molecule. There are local minima along the way that can trap the protein in a metastable state for some time, slowing its progress towards the free energy minimum. As the protein starts to fold, the free energy drops and the number of available conformational states (denoted by the width of the funnel) decreases. The high free energy means that the molecule is unstable, and flops easily between the different conformational states. The high entropy corresponds to there being a large number of possible conformational states-the molecule can take on many different three-dimensional shapes. In this model, the unfolded protein had both high entropy and high free energy. Funnel represents how proteins fold into their native structures that based on concept of minimizing free energy. The native structure has the lowest energy. Each cleft in the photo represents a potentially misfolded protein. Protein must proceed from a high energy/ high entropy state to a low entropy/ low energy state. This energetically favorable folding of the linear polypeptide chain into a stable three-dimensional structure ultimately determines protein function, The macromolecular protein structure is stabilized by the formation of large numbers of weak noncovalent interactions between atoms within the protein. These hydrophobic effects act as a driving force for the protein to assume its three-dimensional shape. This is a more energetically favorable state, so hydrophobic regions do tend to pack together. When hydrophobic regions cluster together, they have less exposed surface area, and fewer water molecules need to be ordered. This is energetically unfavorable, largely because of the decrease in entropy of the water molecules through the restriction of their motion. Because hydrophobic molecules cannot form hydrogen bonds with water, the water that surrounds a hydrophobic region becomes more ordered to satisfy its hydrogen-bonding potential through interactions with other water molecules, forming cage-like structures (Figure 2.24). A hydrophobic effect is due to the tendency of hydrophobic molecules to pack close together away from water. It leads to aquiring a hydration layer where water can have interactions with hydrophillic heads. Going from unfolded to folded- hydrophobic effect plays a major role. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
